Project Objective
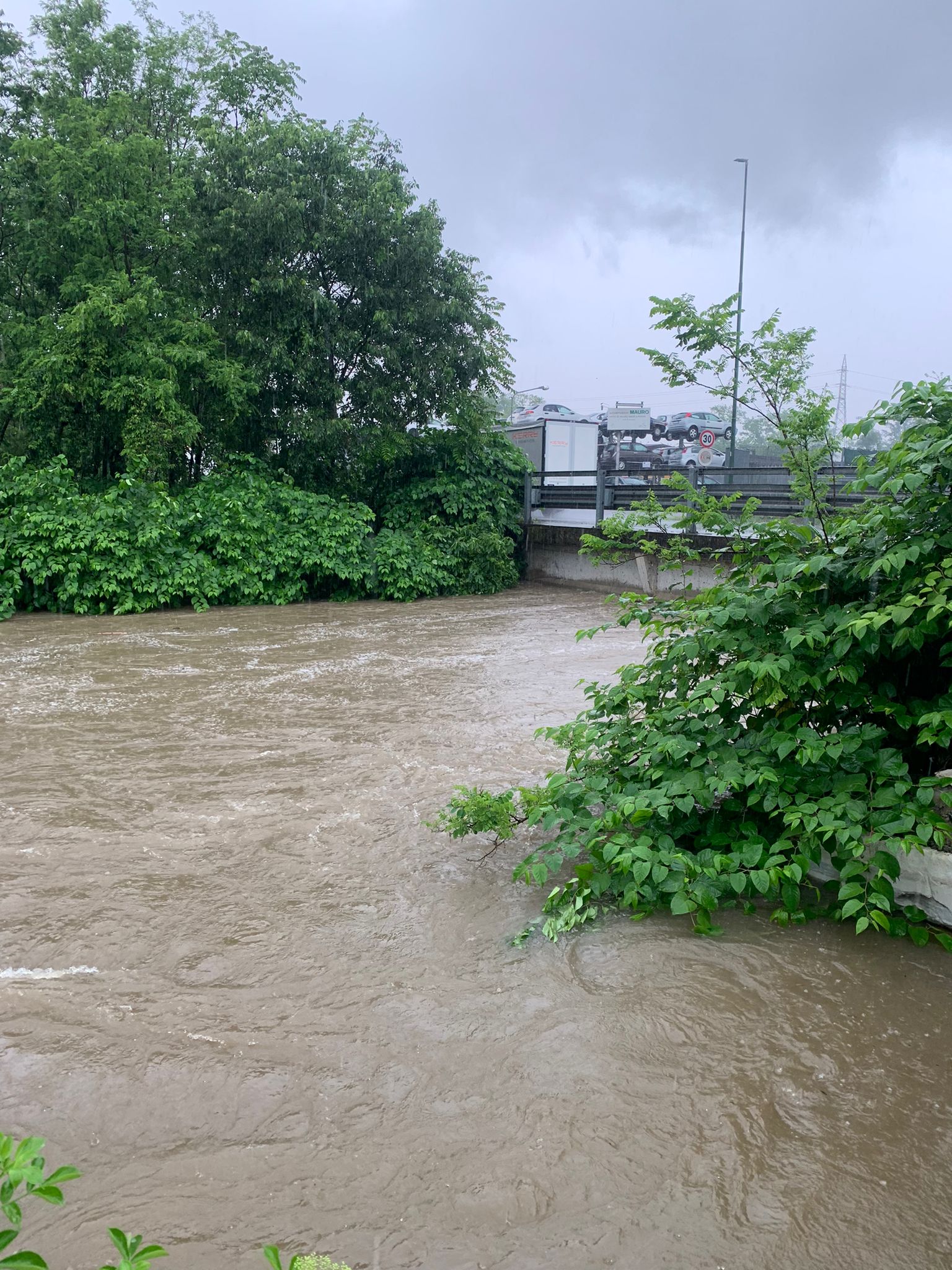
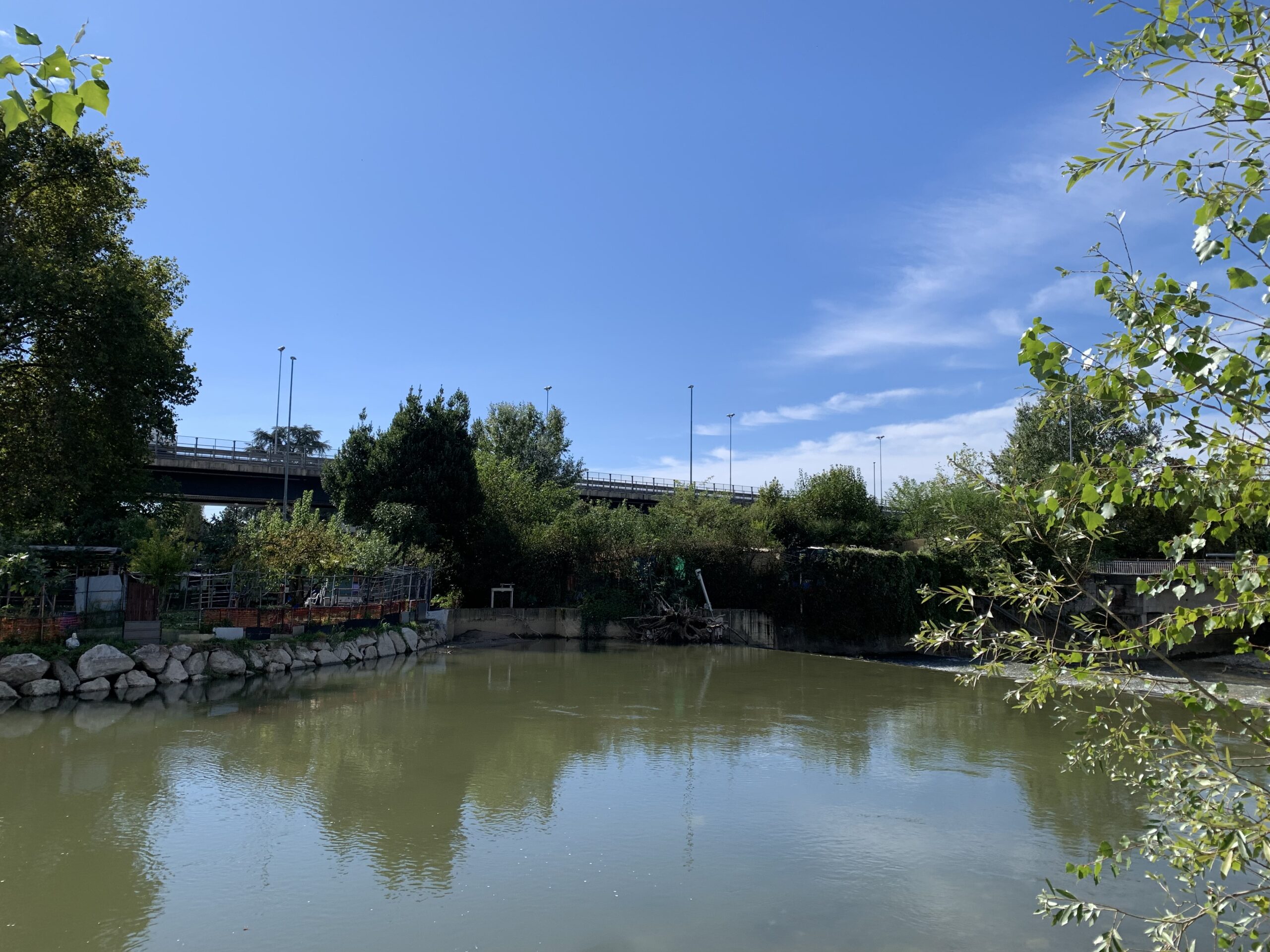
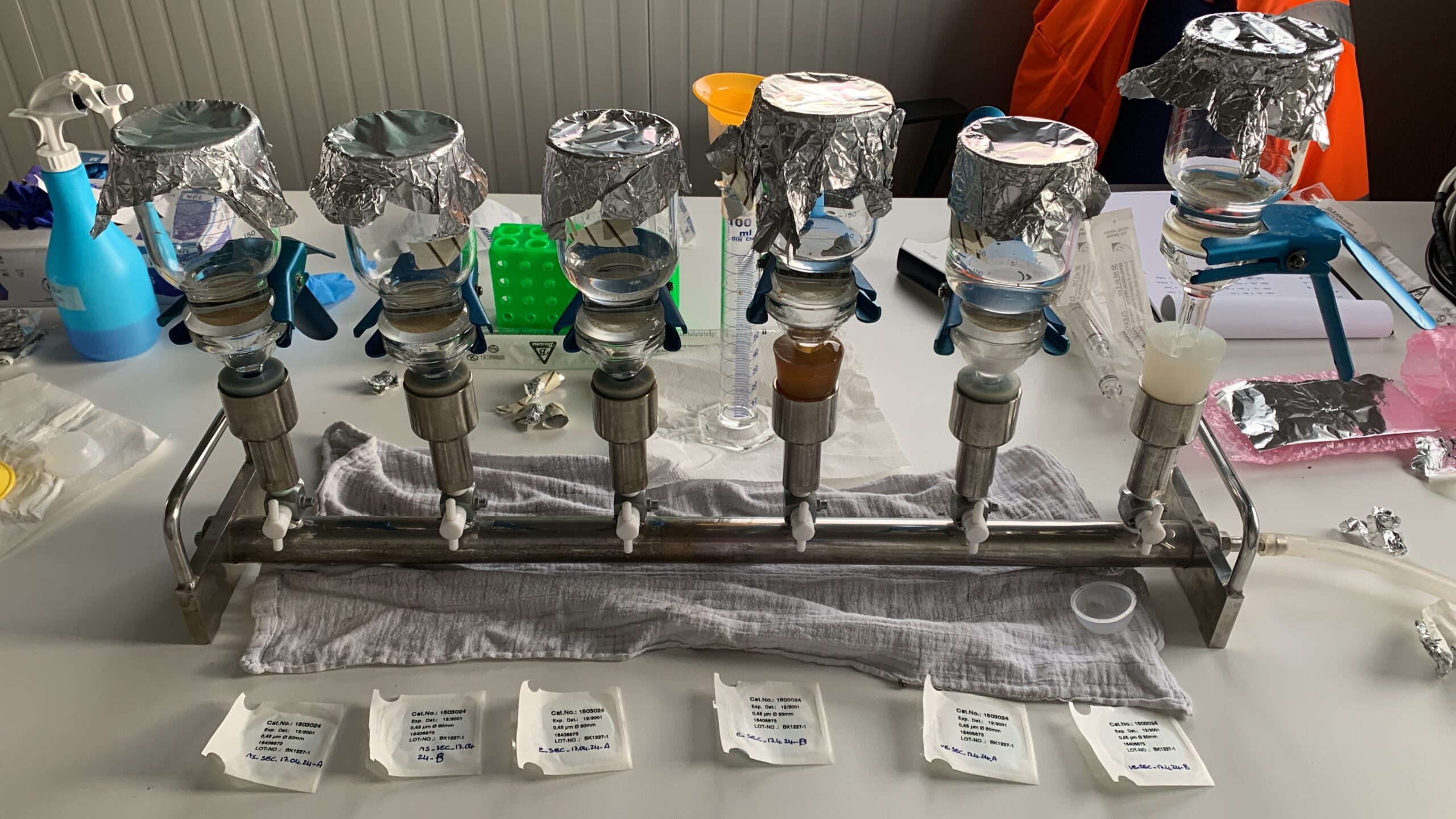
This project aims to investigate the impact of decontamination processes in an urban wastewater treatment plant (WWTP), as well as the potential impact of the wastewater bypass on the release into the environment of bacterial resistome. Specifically, it will assess the presence and concentrations of antibiotic-resistant bacteria (ARB) and antibiotic resistance genes (ARGs) during both dry weather conditions and high-intensity rainfall events, using a combination of culture-dependent and -independent methods.
General description
The discovery of antibiotics for the treatment of infectious diseases represents one of the most important medical achievements of the 20th century. However, their extensive use and misuse have led to a rise in antibiotic-resistant bacteria (ARB), which have acquired antibiotic resistance genes (ARGs) as part of their chromosome and/or plasmids. A wide variety of genes can be responsible for antimicrobial resistance (AMR), and their identification is crucial to understanding the epidemiology and spread of antibiotic-resistant pathogens. ARBs and their related ARGs have been detected in various environmental matrices, including the effluents of urban wastewater treatment plants (UWWTPs). Furthermore, Combined Sewer Overflows (CSOs) may represent a significant pathway for the release of AMR from municipal wastewater into the environment. In a Combined Sewer System (CSS), also known as a wastewater bypass, both wastewater and stormwater are collected in the same pipe. When the volume of sewage in the system exceeds the capacity of the UWWTP, typically during periods of heavy rainfall or snowmelt, the overflow, containing both untreated wastewater and stormwater, is discharged directly into nearby water bodies. Additionally, climate change is expected to increase the frequency and volume of CSOs in certain regions due to shifts in precipitation patterns.
Collaborators
BrianzAcque s.r.l. (Monza, Italy)